QC-Flex
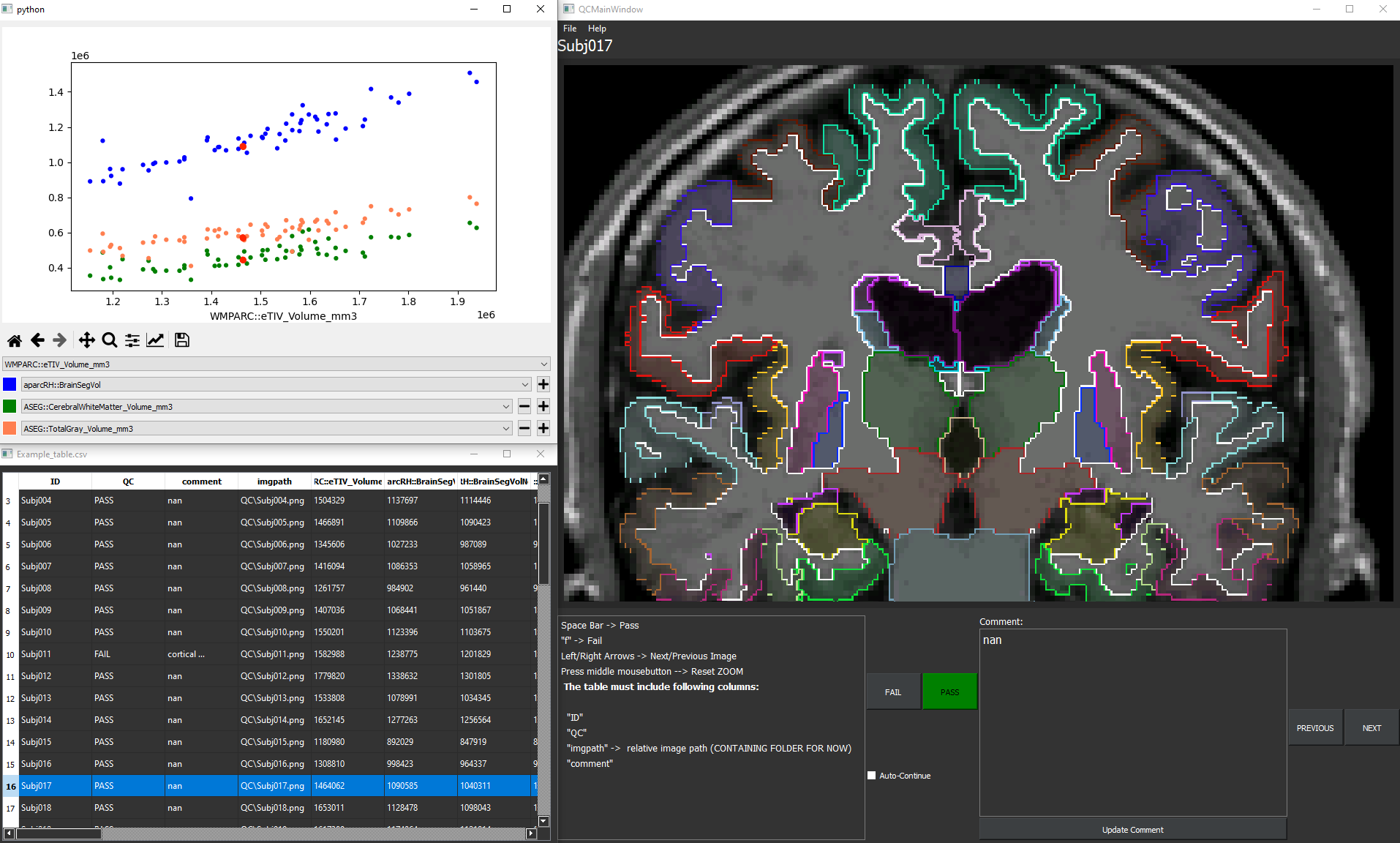
QC-Flex is qulity check framework with flexible application. It can be used for all numeric datasets, which provide a 2D QC image for each entry (subject).
QCFlex: A flexible quality control tool for large MRI cohorts
This work will be presented at the 38rd annual (online) meeting of the ESMRMB 2021 6-9 October 2021
Presentation number: TBA
Abstract ID: 216
DOI: TBA
If you find the provided tool useful for your work, please cite:
Eggenreich S., “QCFlex: A flexible quality control tool for large MRI cohorts”, ESMRMB 38th Annual Scientific Meeting, 7-9 October 2021
Purpose of the software:
The accurate segmentation of brain tissue plays an important role in neuroscience for several purposes. The assessment of global and regional tissue labels, for example, has become a crucial application in several neurodegenerative diseases1,2 and also in normal ageing3, to study subject specific volumetric changes or to perform regional quantitative analyses. Although advanced brain segmentation tools, such as FreeSurfer4, have become more robust, segmentation errors still might occur. Therefore, manual quality assurance is still indispensable, which however becomes difficult and time consuming at a larger scale. Here, we present a tool for fast, explorative and interactive quality assessment optimally suited for larger cohorts, combining the visualization of a segmentation output along with the numeric outcome of the resulting dataset.
Methods
QCFlex was developed under Python 3.9.5 with the Python packages Numpy, Matplotlib, pandas, Pillow, qimage2ndarray. The GUI was built with PyQT5 and can be distributed as an executable. A single table file builds the basis of variables to explore and the location of corresponding image paths.
The interactive image viewer displays a precomputed high resolution 2D image of the resulting segmentations, which can be inspected in detail by zooming into the region of interest. The integrated comment feature allows the annotation of images and storage of those comments in the table file for later use.
The integrated and interactable scatterplots, depicting two selectable outcome-measures (such as brain volume and gray matter volume), allow an instant overview of the data distribution. This feature facilitates the detection of outliers, which can be further inspected by clicking on the datapoint.
Buttons to annotate the images with Pass/Fail further supports the user’s ability to assess the overall quality of the calculations, while still being able to keep track of outliers.
This project was tested with python version 3.9.5, with the requirements being distributed in the requirements.txt file.
In order to quickly setup a development environment, install venv from an installed version of Python 3.9.5 with
Try it out!
Download latest release:
For developers
Install virtualenv from pip
> python3 -m venv <_ENVIRONMENT_NAME_>
On windows activate the installed environment by
> <_ENVIRONMENT_NAME_>\Scripts\activate
and install all dependencies with
> pip install -r requirements.txt
Happy Coding!
Building Release
In order to distribute this project as an executable, run in the root folder of the repository:
> python setup.py bdist
Avoid running the build process in an Anaconda environment to keep the package size as small as possible!

References
- Popescu, V. et al. Grey Matter Atrophy in Multiple Sclerosis: Clinical Interpretation Depends on Choice of Analysis Method. PLoS One 11, e0143942 (2016).
- Brain volumes and their ratios in Alzheimer´s disease on magnetic resonance imaging segmented using Freesurfer 6.0. Psychiatry Research: Neuroimaging 287, 70–74 (2019).
- Dickie, D. A. et al. Variance in brain volume with advancing age: implications for defining the limits of normality. PLoS One 8, e84093 (2013).
- FreeSurfer. Neuroimage 62, 774–781 (2012).
Authors
S. Eggenreich, S. Ropele, L. Pirpamer
Neuroimaging Research Unit - Medical University of Graz, Department of Neurology, Graz, Austria;